A new technique developed by Weill Cornell Medicine investigators that creates an unusually stable form of RNA may illuminate many cellular processes and enable new kinds of gene therapies. A study describing the technique was published April 8 in Nature Biotechnology.
Short strands of RNA, called RNA aptamers, can be very useful tools for scientists. They can bind to proteins and block or control their activity. They can also make it easier for scientists to observe processes inside a cell. But so far, scientists haven’t had much success using these tools in cells from humans or other mammals because the RNA aptamers break apart too quickly and never accumulate to high enough levels to be useful.
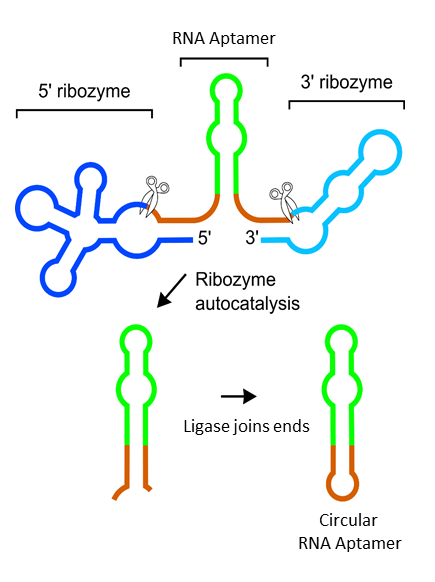
The ribozymes (blue) cleave themselves spontaneously (scissors), leaving the aptamer RNA (green) with chemical ends (brown) that are recognized by the ligase. The ligase then connects the ends of the RNA to create circular RNA. Image courtesy of Samie Jaffrey.
Weill Cornell Medicine investigators found a way to overcome this problem by developing a system called ‘Twister-optimized RNA for durable overexpression,’ or Tornado, that makes circular RNAs, which are more stable. The circular form allows the RNA to escape being broken apart by enzymes that latch on to the ends of linear strands of RNA.
“We now have the possibility of manipulating and probing cellular function using RNA and RNA-based tools, which will allow us to create new types of gene therapies,” said Dr. Samie Jaffrey, the Greenberg-Starr Professor of Pharmacology at Weill Cornell Medicine.
The approach takes advantage of a natural enzymatic reaction that cells use to make transfer RNA (tRNA), a molecule used for protein synthesis. Shortly after tRNA is synthesized in the nucleus, it is cut to remove extra RNA sequences during its initial processing. The tRNA halves are then joined by an enzyme called a ligase.
Jacob Litke, a graduate student in the Tri-Institutional Ph.D. Program in Chemical Biology offered jointly by Weill Cornell Graduate School of Medical Sciences, The Rockefeller University and Memorial Sloan Kettering Cancer Center, wondered if they could take advantage of this ligase to produce circular RNA from RNA aptamers expressed in mammalian cells.
The major challenge was to coax the ligase into acting on RNAs other than tRNA. Normally, the tRNA is cut in a way to produce chemically unique ends on the RNA fragments that are recognized by the ligase. To create similar ends on RNA aptamers expressed in the cell, the team sandwiched the RNA between two sequences of bacterial RNA called ribozymes. The ribozymes then cut themselves automatically, leaving behind the desired chemically unique ends at the site of cleavage. The ligase then connects the ends of the RNA aptamer to create a circular RNA aptamer.
The team found that these circular RNA aptamers don’t break down as quickly and accumulate to levels that are 200 times higher than linear aptamers, and about 50 times higher than circular RNA aptamers produced using other methods, Dr. Jaffrey said. This results in enough RNA aptamers to potentially bind to every one of their protein targets and may permit their therapeutic use.
“It’s a dramatic improvement,” he said.
Dr. Jaffrey and Litke also showed that some circular RNAs can act as sensors to report on cellular processes with fluorescence, enabling researchers to watch the metabolism of living mammalian cells in the laboratory, which was previously only possible in bacterial cells. They also conducted several experiments to test whether the circular RNAs harm cells or trigger immune reactions and found they do not.
“With the Tornado expression system, we can stably express RNAs in mammalian cells,” Litke said, “and the whole world of RNA-based tools can now be used to manipulate and probe cellular function.”
Dr. Jaffrey is the founder and scientific advisor to Lucerna Technologies and Gotham Therapeutics, and holds equity interests in these companies. In addition, Lucerna Technologies has a license related to technology described in this story.

